 ]]>
]]>#!/bin/bash
NCPUS=4 # threads per micrograph
JOBS=6 # micrographs in parallel
CTF=/data/csun/softwares/ctf/ctffind3_mp.exe
run_one () {
mrc="$1"
log="${mrc%.mrc}_ctffind3.log"
[[ -f "$log" ]] && { echo "Skipping $mrc"; return 0; }
export NCPUS="$NCPUS"
"$CTF" << EOF > "$log" 2>&1
$mrc
${mrc%.mrc}_ctf.mrc
2.7,300.0,0.1,28902,2.5
512,50,5,50000,5000,500,900
EOF
echo "Done $mrc"
}
export -f run_one
export NCPUS CTF
parallel -j "$JOBS" run_one ::: *patch_aligned.mrc
 ]]>
]]>e2iminfo.py -H micrograph.mrc
e2proc2d.py input.mrc output.tif
e2proc2d.py *mrc @tif
e2proc3d.py input.hdf output.hdf --clip=140,140,140 --scale=2.8
e2project3d.py map.mrc --outfile=proj.mrcs --orientgen rand:n=10:phitoo=1:trans=4
e2proc3d.py input.mrc output.mrc --process filter.fourier:curres=3.5:targetres=7:dampamp=1:randomizephase=0
e2proc3d.py input.mrc output.mrc --process filter.autosharpening:cref=1:mapres=2.5:lpfile=fsc.txt.fit
e2proc3d.py input.mrc output.mrc --calcsf output_sf.txt
e2proc3d.py input.mrc output.mrc --apix 1.08 --setsf sf.txt
e2proc3d.py input.mrc output.mrc --apix 1.08 --matchto other.mrc
e2project3d.py map.mrc --outfile=projections.mrcs --orientgen random:n=1000:phitoo=1:inc_mirror=1:trans=10
e2proc3d.py input.mrc output.mrc --rot az,alt,phi
e2proc3d.py input.mrc output.mrc --rotspin n1,n2,n3,angle
e2proc3d.py input.mrc output.mrc --oct4fto3f
e2proc3d.py input.mrc output.mrc --rot 45,54.7356,0
e2proc3d.py input.mrc output.mrc --process xform.flip:axis=o
e2proc3d.py input.mrc output.mrc --process xform.flip:axis=y
e2proc3d.py input.mrc output.mrc --process mask.file:file=mask.mrc
e2proc3d.py input.mrc output.mrc --process mask.cylinder3d:innerradius=10A:outerradius=20A:length=60A:masksoft=10A
e2proc3d.py :96:96:96:1 output.mrc --apix 1 --process mask.cylinder3d:innerradius=10A:outerradius=20A:length=60A:masksoft=10A
relion_mask_create --denovo --box_size 256 --inner_radius 0 --outer_radius 50 --angpix 1.6 --o mask.mrc
e2proc3d.py input.mrc output.mrc --process mask.auto3d:nshells=5:nshellsgauss=5:thresh=3
e2proc3d.py input.mrc ouput.mrc --process xform.helical:apix=1.1:az0=22:z0=37.2:cdsym=c6:verbose=1:rstep=1
e2proc3d.py input.mrc ouput.mrc --process xform.helical:apix=1.1:az0=22:z0=37.2:cdsym=c6:verbose=1
Fourier Shell Correlation (FSC) is the standard method used in the cryo-electron microscopy (cryo-EM) field to estimate the resolution of reconstructed 3D maps, serving as a key indicator of overall map quality. FSC is computed as the cross-correlation coefficient between two independently reconstructed “half maps” as a function of spatial frequency. At each resolution shell, FSC quantifies how well the two maps agree, with higher values indicating better reproducibility and signal.
However, a single FSC value per shell assumes uniformity in signal quality across all voxels within that shell. In practice, this assumption oversimplifies the reality: voxel intensities vary substantially across different spatial locations, even within the same frequency band. This spatial heterogeneity can arise from anisotropic sampling, preferred particle orientations, or local disorder in the structure.
To better capture these local variations, this web application computes FSC not just as an average, but includes statistical confidence intervals around each FSC value. This web app allows you to visualize confidence intervals around the FSC values using three different statistical approaches. You can select your preferred method from the dropdown menu:
This approach assumes the Fisher z-transformed FSC values follow a normal distribution, allowing us to compute analytical confidence bounds. Formula:
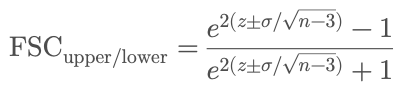
Where:
z = 0.5 × log((1 + FSC) / (1 - FSC))
is the Fisher z-transform of the FSC
n is the number of voxels in the resolution shell
σ (sigma) is a user-defined z-score (e.g., 1.96 for 95% confidence, 3 for 99.7%)
By allowing you to set the sigma value, this app gives you flexible control over the statistical stringency of the FSC envelope, helping you better interpret the reliability of FSC-based resolution estimates.
This non-parametric approach resamples voxel pairs within each shell to build a distribution of FSC values. Confidence bounds are calculated from percentiles of this distribution.
This method estimates FSC uncertainty analytically using a published variance formula that accounts for the number of voxels and the observed FSC value. Confidence bounds are derived as:
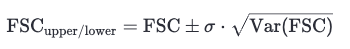
Where Var(FSC) is computed using:
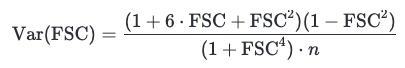
frequency = shell_index / (box_size * apix) # spatial frequency in 1/Å
resolution = 1 / frequency
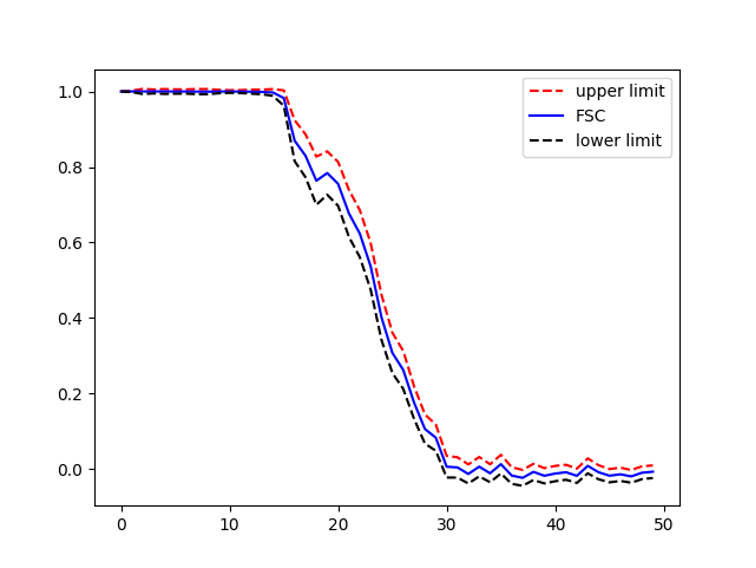
Example FSC error bar for EMPIAR 10084 Cryo-EM structure of haemoglobin at 3.2 Å determined with the Volta phase plate
Penczek PA. Resolution measures in molecular electron microscopy. Methods Enzymol. 2010; 482:73–100
PMID: 20888958 | PMCID: PMC3165049
Cardone G, Heymann JB, Steven AC. One number does not fit all: mapping local variations in resolution in cryo-EM reconstructions. J Struct Biol. 2013; 184(2):226–236
PMID: 23954653 | PMCID: PMC3837392
 ]]>
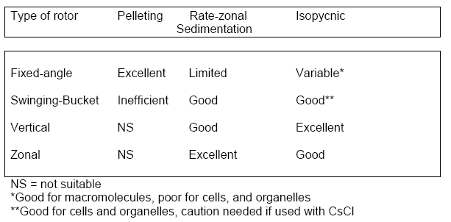
]]>Density gradient centrifugation can be further divided into isopycnic centrifugation and rate-zonal centrifugation.
Isopycnic centrifugation separates particles only based on their buoyant density. It uses a continuous density gradient, which can be either preformed or self-forming (e.g., using CsCl or OptiPrep). In self-forming gradients, the sample is mixed with the gradient material, and the gradient forms during centrifugation. This process typically requires long run times—often overnight—to allow the gradient and particle separation to equilibrate.
Rate-zonal centrifugation separates particles based on their size and sedimentation rate. It often uses a stepwise gradient—a series of discrete layers with increasing density (e.g., 10%, 20%, 30%, 40%). The sample is layered on top of the lightest density layer. This method is widely used for virus purification and typically requires shorter centrifugation times (e.g., one to two hours). However, if spun too long, all particles may eventually pellet, reducing separation resolution.
Sometimes one has to run isopycnic and rate-zonal centrifugation in tandem to enhance purity.
⚠️ Care must be taken to avoid “point loads” caused by spinning CsCl or other dense gradient materials that can precipitate.
The centrifugation time is given by:
\[t = \frac{k}{s'}\]The k factor is a measure of the pelleting efficiency of the rotor.
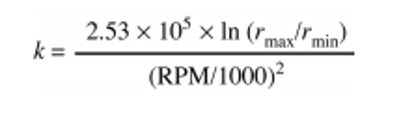
As the k factor decreases, rotor efficiency increases.
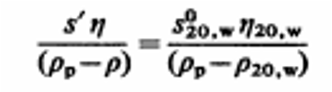 Biochem. J. (1976) 159, 259-265
Biochem. J. (1976) 159, 259-265
Where:
At 4°C in 20% sucrose, the sedimentation coefficient of Tulane virus:
Calculation: \(t = \frac{k}{s'} = \frac{72.4}{41.87} \approx 1.73\ \text{h} \approx 2\ \text{hours}\) —
Note: The sedimentation coefficient is influenced by the viscosity and density of the sucrose solution. Viscosity is dependent on both concentration and temperature.
]]>Check out the Practical Techniques for Centrifugal Separations for more details.
The tutorial commands result in error messages.
git clone https://github.com/czimaginginstitute/AreTomo3.git
cd AreTomo3
make -f makefile11 CUDAHOME=/usr/local/cuda \
CXXFLAGS="-I/usr/local/cuda/include" \
LDFLAGS="-L/usr/local/cuda/lib64"
MatrixScreener → Matrix MAPS + CLEM⚠️ Do not remove the hose before the temperature reaches 60°C.
Removing the hose prematurely can cause damage due to freezing.
\\AS-SPC\SharedData\Chen\20241106⚠️ Repeat Steps 5–8 for Grid 2.
⚠️ Do not exceed 10 seconds. A thick GIS layer will hinder milling efficiency.
Grid1_afterGIS-XXX.tif.D:\AuToTEM Cryo\Projects).A thin lamella should appear black in SEM.
File → Save screenshot as → Save grid image with lamella positions.L2_i.tifL2_s_2kV.tifL2_1200x_2kV.tif (for LM correlation)⚠️ This is important to prevent ion beam waste.
Clone the GitHub repository and follow the instructions in the README to install dependencies and set up the Python environment:
To specify which GPU to use, prepend your MemBrain command with the CUDA_VISIBLE_DEVICES environment variable. This variable tells PyTorch which GPU(s) are visible to the program. You can run multiple MemBrain commands in parallel on different GPUs to process several tomograms simultaneously.
CUDA_VISIBLE_DEVICES=1 membrain segment --tomogram-path lamella4_ts_001_full_rec.mrc --ckpt-path /home/csun/Membrain-model/MemBrain_seg_v10_alpha.ckpt --store-connected-components